Note
Click here to download the full example code or to run this example in your browser via Binder
Fitting#
Learn how the model, dataset and fit Gammapy classes work together in a detailed modeling and fitting use-case.
Prerequisites#
Knowledge of spectral analysis to produce 1D On-Off datasets, see the Spectral analysis tutorial.
Reading of pre-computed datasets see e.g. Multi instrument joint 3D and 1D analysis tutorial.
General knowledge on statistics and optimization methods
Proposed approach#
This is a hands-on tutorial to modeling, showing how to do
perform a Fit in gammapy. The emphasis here is on interfacing the
Fit class and inspecting the errors. To see an analysis example of
how datasets and models interact, see the Modelling tutorial.
As an example, in this notebook, we are going to work with HESS data of the Crab Nebula and show in
particular how to :
perform a spectral analysis
use different fitting backends
access covariance matrix information and parameter errors
compute likelihood profile - compute confidence contours
See also: Models and Modeling and Fitting (DL4 to DL5).
The setup#
from itertools import combinations
import numpy as np
from astropy import units as u
import matplotlib.pyplot as plt
from IPython.display import display
from gammapy.datasets import Datasets, SpectrumDatasetOnOff
from gammapy.modeling import Fit
from gammapy.modeling.models import LogParabolaSpectralModel, SkyModel
Check setup#
from gammapy.utils.check import check_tutorials_setup
from gammapy.visualization.utils import plot_contour_line
check_tutorials_setup()
System:
python_executable : /home/runner/work/gammapy-docs/gammapy-docs/gammapy/.tox/build_docs/bin/python
python_version : 3.9.15
machine : x86_64
system : Linux
Gammapy package:
version : 1.0
path : /home/runner/work/gammapy-docs/gammapy-docs/gammapy/.tox/build_docs/lib/python3.9/site-packages/gammapy
Other packages:
numpy : 1.23.4
scipy : 1.9.3
astropy : 5.1.1
regions : 0.7
click : 8.1.3
yaml : 6.0
IPython : 8.6.0
jupyterlab : not installed
matplotlib : 3.6.2
pandas : not installed
healpy : 1.16.1
iminuit : 2.17.0
sherpa : 4.15.0
naima : 0.10.0
emcee : 3.1.3
corner : 2.2.1
Gammapy environment variables:
GAMMAPY_DATA : /home/runner/work/gammapy-docs/gammapy-docs/gammapy-datasets/1.0
Model and dataset#
First we define the source model, here we need only a spectral model for which we choose a log-parabola
crab_spectrum = LogParabolaSpectralModel(
amplitude=1e-11 / u.cm**2 / u.s / u.TeV,
reference=1 * u.TeV,
alpha=2.3,
beta=0.2,
)
crab_spectrum.alpha.max = 3
crab_spectrum.alpha.min = 1
crab_model = SkyModel(spectral_model=crab_spectrum, name="crab")
The data and background are read from pre-computed ON/OFF datasets of HESS observations, for simplicity we stack them together. Then we set the model and fit range to the resulting dataset.
datasets = []
for obs_id in [23523, 23526]:
dataset = SpectrumDatasetOnOff.read(
f"$GAMMAPY_DATA/joint-crab/spectra/hess/pha_obs{obs_id}.fits"
)
datasets.append(dataset)
dataset_hess = Datasets(datasets).stack_reduce(name="HESS")
datasets = Datasets(datasets=[dataset_hess])
# Set model and fit range
dataset_hess.models = crab_model
e_min = 0.66 * u.TeV
e_max = 30 * u.TeV
dataset_hess.mask_fit = dataset_hess.counts.geom.energy_mask(e_min, e_max)
Fitting options#
First let’s create a Fit instance:
scipy_opts = {
"method": "L-BFGS-B",
"options": {"ftol": 1e-4, "gtol": 1e-05},
"backend": "scipy",
}
fit_scipy = Fit(store_trace=True, optimize_opts=scipy_opts)
By default the fit is performed using MINUIT, you can select alternative
optimizers and set their option using the optimize_opts argument of
the Fit.run() method. In addition we have specified to store the
trace of parameter values of the fit.
Note that, for now, covaraince matrix and errors are computed only for the fitting with MINUIT. However depending on the problem other optimizers can better perform, so sometimes it can be useful to run a pre-fit with alternative optimization methods.
result_scipy = fit_scipy.run(datasets)
sherpa_opts = {"method": "simplex", "ftol": 1e-3, "maxfev": int(1e4)}
fit_sherpa = Fit(store_trace=True, backend="sherpa", optimize_opts=sherpa_opts)
results_simplex = fit_sherpa.run(datasets)
For the “minuit” backend see https://iminuit.readthedocs.io/en/latest/reference.html for a detailed description of the available options. If there is an entry ‘migrad_opts’, those options will be passed to iminuit.Minuit.migrad. Additionally you can set the fit tolerance using the tol option. The minimization will stop when the estimated distance to the minimum is less than 0.001*tol (by default tol=0.1). The strategy option change the speed and accuracy of the optimizer: 0 fast, 1 default, 2 slow but accurate. If you want more reliable error estimates, you should run the final fit with strategy 2.
fit = Fit(store_trace=True)
minuit_opts = {"tol": 0.001, "strategy": 1}
fit.backend = "minuit"
fit.optimize_opts = minuit_opts
result_minuit = fit.run(datasets)
Fit quality assessment#
There are various ways to check the convergence and quality of a fit. Among them:
Refer to the automatically-generated results dictionary:
print(result_scipy)
OptimizeResult
backend : scipy
method : L-BFGS-B
success : True
message : CONVERGENCE: REL_REDUCTION_OF_F_<=_FACTR*EPSMCH
nfev : 60
total stat : 30.35
CovarianceResult
backend : minuit
method : hesse
success : True
message : Hesse terminated successfully.
print(results_simplex)
OptimizeResult
backend : sherpa
method : simplex
success : True
message : Optimization terminated successfully
nfev : 135
total stat : 30.35
print(result_minuit)
OptimizeResult
backend : minuit
method : migrad
success : True
message : Optimization terminated successfully.
nfev : 37
total stat : 30.35
CovarianceResult
backend : minuit
method : hesse
success : True
message : Hesse terminated successfully.
If the fit is performed with minuit you can print detailed informations to check the convergence
print(fit.minuit)
┌─────────────────────────────────────────────────────────────────────────┐
│ Migrad │
├──────────────────────────────────┬──────────────────────────────────────┤
│ FCN = 30.35 │ Nfcn = 37 │
│ EDM = 3.42e-08 (Goal: 2e-06) │ time = 0.1 sec │
├──────────────────────────────────┼──────────────────────────────────────┤
│ Valid Minimum │ No Parameters at limit │
├──────────────────────────────────┼──────────────────────────────────────┤
│ Below EDM threshold (goal x 10) │ Below call limit │
├───────────────┬──────────────────┼───────────┬─────────────┬────────────┤
│ Covariance │ Hesse ok │ Accurate │ Pos. def. │ Not forced │
└───────────────┴──────────────────┴───────────┴─────────────┴────────────┘
┌───┬───────────────────┬───────────┬───────────┬────────────┬────────────┬─────────┬─────────┬───────┐
│ │ Name │ Value │ Hesse Err │ Minos Err- │ Minos Err+ │ Limit- │ Limit+ │ Fixed │
├───┼───────────────────┼───────────┼───────────┼────────────┼────────────┼─────────┼─────────┼───────┤
│ 0 │ par_000_amplitude │ 3.8 │ 0.4 │ │ │ │ │ │
│ 1 │ par_001_alpha │ 2.20 │ 0.26 │ │ │ 1 │ 3 │ │
│ 2 │ par_002_beta │ 2.3 │ 1.4 │ │ │ │ │ │
└───┴───────────────────┴───────────┴───────────┴────────────┴────────────┴─────────┴─────────┴───────┘
┌───────────────────┬───────────────────────────────────────────────────────┐
│ │ par_000_amplitude par_001_alpha par_002_beta │
├───────────────────┼───────────────────────────────────────────────────────┤
│ par_000_amplitude │ 0.126 0.0455 -0.117 │
│ par_001_alpha │ 0.0455 0.0689 -0.331 │
│ par_002_beta │ -0.117 -0.331 1.95 │
└───────────────────┴───────────────────────────────────────────────────────┘
Check the trace of the fit e.g. in case the fit did not converge properly
display(result_minuit.trace)
total_stat crab.spectral.amplitude ... crab.spectral.beta
------------------ ----------------------- ... -------------------
30.34953055035606 3.812242570863471e-11 ... 0.22648273120014317
30.349724590233844 3.815793905090241e-11 ... 0.22648273120014317
30.34971174985044 3.808691236636701e-11 ... 0.22648273120014317
30.349539326678645 3.8129513940414386e-11 ... 0.22648273120014317
30.349536722649045 3.811533747685503e-11 ... 0.22648273120014317
30.350508396085097 3.812242570863471e-11 ... 0.22648273120014317
30.35054344544943 3.812242570863471e-11 ... 0.22648273120014317
30.349538987558546 3.812242570863471e-11 ... 0.22648273120014317
30.349542020536582 3.812242570863471e-11 ... 0.22648273120014317
30.350294833727336 3.812242570863471e-11 ... 0.227889227119385
... ... ... ...
30.349537804469417 3.812216975010786e-11 ... 0.226626246441772
30.349538077833646 3.812216975010786e-11 ... 0.22635142719749335
30.349530774659502 3.812358739672099e-11 ... 0.2264888368196327
30.349530758085038 3.812075210349472e-11 ... 0.2264888368196327
30.349530807483546 3.812216975010786e-11 ... 0.2264888368196327
30.349530725219488 3.812216975010786e-11 ... 0.2264888368196327
30.349530739447403 3.812216975010786e-11 ... 0.22651631874406056
30.349530793246174 3.812216975010786e-11 ... 0.2264613548952048
30.349535814960312 3.8129257983173524e-11 ... 0.2264888368196327
30.349537158863242 3.8129257983173524e-11 ... 0.226626246441772
30.349559366906988 3.812216975010786e-11 ... 0.226626246441772
Length = 37 rows
The fitted models are copied on the FitResult object.
They can be inspected to check that the fitted values and errors
for all parameters are reasonable, and no fitted parameter value is “too close”
- or even outside - its allowed min-max range
display(result_minuit.models.to_parameters_table())
model type name value ... max frozen is_norm link
----- -------- --------- ---------- ... --------- ------ ------- ----
crab spectral amplitude 3.8122e-11 ... nan False True
crab spectral reference 1.0000e+00 ... nan True False
crab spectral alpha 2.1958e+00 ... 3.000e+00 False False
crab spectral beta 2.2649e-01 ... nan False False
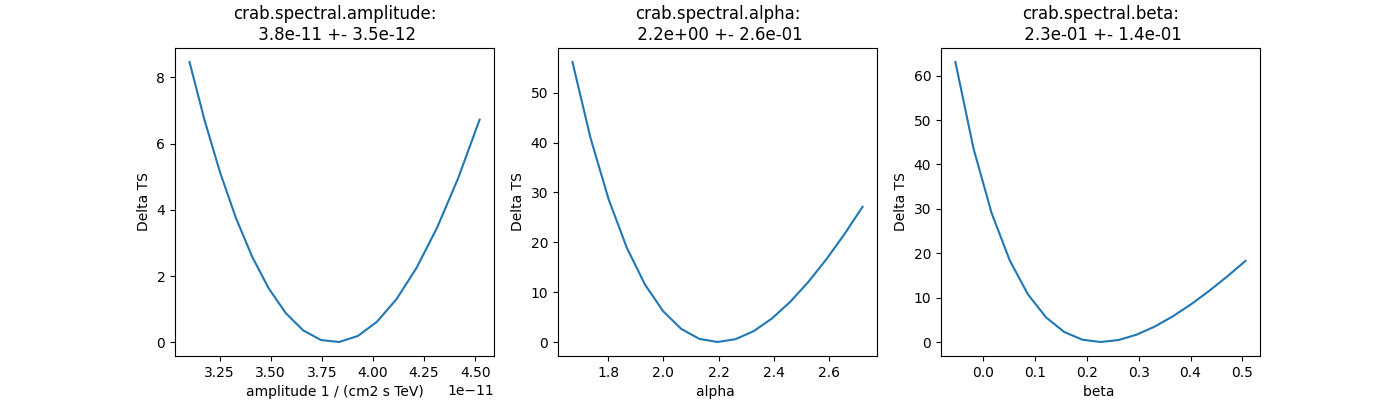
Plot fit statistic profiles for all fitted parameters, using
stat_profile. For a good fit and error
estimate each profile should be parabolic. The specification for each
fit statistic profile can be changed on the
Parameter object, which has scan_min,
scan_max, scan_n_values and scan_n_sigma attributes.
total_stat = result_minuit.total_stat
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=(14, 4))
for ax, par in zip(axes, datasets.parameters.free_parameters):
par.scan_n_values = 17
idx = datasets.parameters.index(par)
name = datasets.models.parameters_unique_names[idx]
profile = fit.stat_profile(datasets=datasets, parameter=par)
ax.plot(profile[f"{name}_scan"], profile["stat_scan"] - total_stat)
ax.set_xlabel(f"{par.name} {par.unit}")
ax.set_ylabel("Delta TS")
ax.set_title(f"{name}:\n {par.value:.1e} +- {par.error:.1e}")
Inspect model residuals. Those can always be accessed using
residuals(). For more details, we refer here to the dedicated
3D detailed analysis (for MapDataset fitting) and
Spectral analysis (for SpectrumDataset fitting).
Covariance and parameters errors#
After the fit the covariance matrix is attached to the models copy
stored on the FitResult object.
You can access it directly with:
print(result_minuit.models.covariance)
[[ 1.25743554e-23 0.00000000e+00 4.54676856e-13 -1.17016492e-13]
[ 0.00000000e+00 0.00000000e+00 0.00000000e+00 0.00000000e+00]
[ 4.54676856e-13 0.00000000e+00 6.89492210e-02 -3.31139115e-02]
[-1.17016492e-13 0.00000000e+00 -3.31139115e-02 1.95024569e-02]]
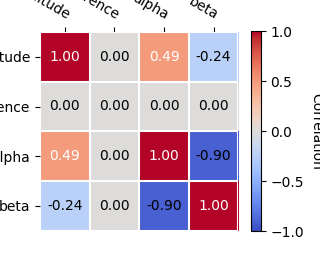
And you can plot the total parameter correlation as well:
result_minuit.models.covariance.plot_correlation()
# The covariance information is also propagated to the individual models
# Therefore, one can also get the error on a specific parameter by directly
# accessing the `~gammapy.modeling.Parameter.error` attribute:
#
print(crab_model.spectral_model.alpha.error)
0.2625818368198403
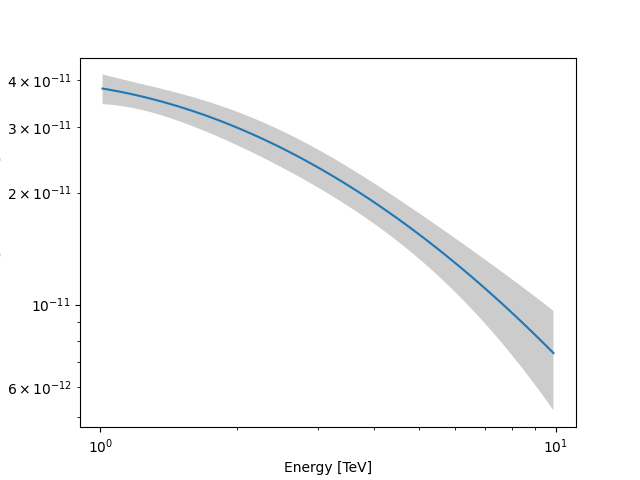
As an example, this step is needed to produce a butterfly plot showing the envelope of the model taking into account parameter uncertainties.
plt.figure()
energy_bounds = [1, 10] * u.TeV
crab_spectrum.plot(energy_bounds=energy_bounds, energy_power=2)
ax = crab_spectrum.plot_error(energy_bounds=energy_bounds, energy_power=2)
Confidence contours#
In most studies, one wishes to estimate parameters distribution using observed sample data. A 1-dimensional confidence interval gives an estimated range of values which is likely to include an unknown parameter. A confidence contour is a 2-dimensional generalization of a confidence interval, often represented as an ellipsoid around the best-fit value.
Gammapy offers two ways of computing confidence contours, in the
dedicated methods minos_contour and stat_profile. In
the following sections we will describe them.
An important point to keep in mind is: what does a :math:`Nsigma` confidence contour really mean? The answer is it represents the points of the parameter space for which the model likelihood is \(N\sigma\) above the minimum. But one always has to keep in mind that 1 standard deviation in two dimensions has a smaller coverage probability than 68%, and similarly for all other levels. In particular, in 2-dimensions the probability enclosed by the \(N\sigma\) confidence contour is \(P(N)=1-e^{-N^2/2}\).
Computing contours using stat_contour#
After the fit, MINUIT offers the possibility to compute the confidence
confours. gammapy provides an interface to this functionality through
the Fit object using the stat_contour method. Here we defined a
function to automate the contour production for the different
parameter and confidence levels (expressed in terms of sigma):
def make_contours(fit, datasets, result, npoints, sigmas):
cts_sigma = []
for sigma in sigmas:
contours = dict()
for par_1, par_2 in combinations(["alpha", "beta", "amplitude"], r=2):
idx1, idx2 = datasets.parameters.index(par_1), datasets.parameters.index(
par_2
)
name1 = datasets.models.parameters_unique_names[idx1]
name2 = datasets.models.parameters_unique_names[idx2]
contour = fit.stat_contour(
datasets=datasets,
x=datasets.parameters[par_1],
y=datasets.parameters[par_2],
numpoints=npoints,
sigma=sigma,
)
contours[f"contour_{par_1}_{par_2}"] = {
par_1: contour[name1].tolist(),
par_2: contour[name2].tolist(),
}
cts_sigma.append(contours)
return cts_sigma
Now we can compute few contours.
Then we prepare some aliases and annotations in order to make the plotting nicer.
pars = {
"phi": r"$\phi_0 \,/\,(10^{-11}\,{\rm TeV}^{-1} \, {\rm cm}^{-2} {\rm s}^{-1})$",
"alpha": r"$\alpha$",
"beta": r"$\beta$",
}
panels = [
{
"x": "alpha",
"y": "phi",
"cx": (lambda ct: ct["contour_alpha_amplitude"]["alpha"]),
"cy": (lambda ct: np.array(1e11) * ct["contour_alpha_amplitude"]["amplitude"]),
},
{
"x": "beta",
"y": "phi",
"cx": (lambda ct: ct["contour_beta_amplitude"]["beta"]),
"cy": (lambda ct: np.array(1e11) * ct["contour_beta_amplitude"]["amplitude"]),
},
{
"x": "alpha",
"y": "beta",
"cx": (lambda ct: ct["contour_alpha_beta"]["alpha"]),
"cy": (lambda ct: ct["contour_alpha_beta"]["beta"]),
},
]
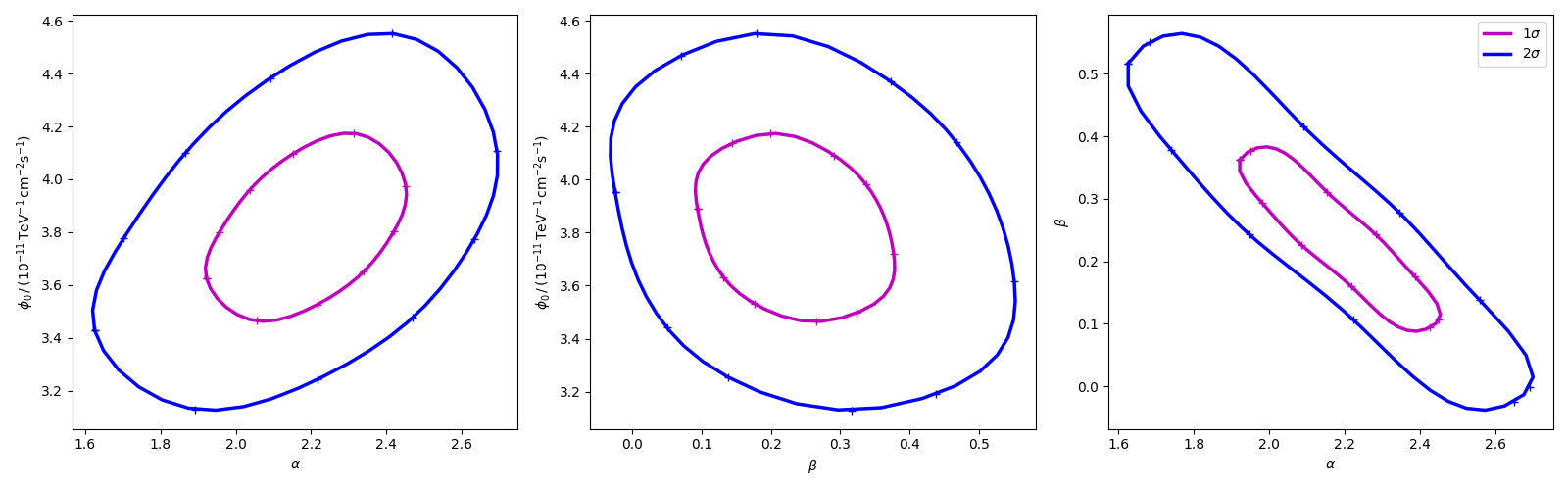
Finally we produce the confidence contours figures.
fig, axes = plt.subplots(1, 3, figsize=(16, 5))
colors = ["m", "b", "c"]
for p, ax in zip(panels, axes):
xlabel = pars[p["x"]]
ylabel = pars[p["y"]]
for ks in range(len(cts_sigma)):
plot_contour_line(
ax,
p["cx"](cts_sigma[ks]),
p["cy"](cts_sigma[ks]),
lw=2.5,
color=colors[ks],
label=f"{sigmas[ks]}" + r"$\sigma$",
)
ax.set_xlabel(xlabel)
ax.set_ylabel(ylabel)
plt.legend()
plt.tight_layout()
Computing contours using stat_surface#
This alternative method for the computation of confidence contours,
although more time consuming than minos_contour(), is expected
to be more stable. It consists of a generalization of
stat_profile() to a 2-dimensional parameter space. The algorithm
is very simple: - First, passing two arrays of parameters values, a
2-dimensional discrete parameter space is defined; - For each node of
the parameter space, the two parameters of interest are frozen. This
way, a likelihood value (\(-2\mathrm{ln}\,\mathcal{L}\), actually)
is computed, by either freezing (default) or fitting all nuisance
parameters; - Finally, a 2-dimensional surface of
\(-2\mathrm{ln}(\mathcal{L})\) values is returned. Using that
surface, one can easily compute a surface of
\(TS = -2\Delta\mathrm{ln}(\mathcal{L})\) and compute confidence
contours.
Let’s see it step by step.
First of all, we can notice that this method is “backend-agnostic”, meaning that it can be run with MINUIT, sherpa or scipy as fitting tools. Here we will stick with MINUIT, which is the default choice:
As an example, we can compute the confidence contour for the alpha
and beta parameters of the dataset_hess. Here we define the
parameter space:
result = result_minuit
par_alpha = datasets.parameters["alpha"]
par_beta = datasets.parameters["beta"]
par_alpha.scan_values = np.linspace(1.55, 2.7, 20)
par_beta.scan_values = np.linspace(-0.05, 0.55, 20)
Then we run the algorithm, by choosing reoptimize=False for the sake
of time saving. In real life applications, we strongly recommend to use
reoptimize=True, so that all free nuisance parameters will be fit at
each grid node. This is the correct way, statistically speaking, of
computing confidence contours, but is expected to be time consuming.
fit = Fit(backend="minuit", optimize_opts={"print_level": 0})
stat_surface = fit.stat_surface(
datasets=datasets,
x=par_alpha,
y=par_beta,
reoptimize=False,
)
In order to easily inspect the results, we can convert the \(-2\mathrm{ln}(\mathcal{L})\) surface to a surface of statistical significance (in units of Gaussian standard deviations from the surface minimum):
# Compute TS
TS = stat_surface["stat_scan"] - result.total_stat
# Compute the corresponding statistical significance surface
stat_surface = np.sqrt(TS.T)
Notice that, as explained before, \(1\sigma\) contour obtained this way will not contain 68% of the probability, but rather
# Compute the corresponding statistical significance surface
# p_value = 1 - st.chi2(df=1).cdf(TS)
# gaussian_sigmas = st.norm.isf(p_value / 2).T
Finally, we can plot the surface values together with contours:
fig, ax = plt.subplots(figsize=(8, 6))
x_values = par_alpha.scan_values
y_values = par_beta.scan_values
# plot surface
im = ax.pcolormesh(x_values, y_values, stat_surface, shading="auto")
fig.colorbar(im, label="sqrt(TS)")
ax.set_xlabel(f"{par_alpha.name}")
ax.set_ylabel(f"{par_beta.name}")
# We choose to plot 1 and 2 sigma confidence contours
levels = [1, 2]
contours = ax.contour(x_values, y_values, stat_surface, levels=levels, colors="white")
ax.clabel(contours, fmt="%.0f $\\sigma$", inline=3, fontsize=15)
plt.show()
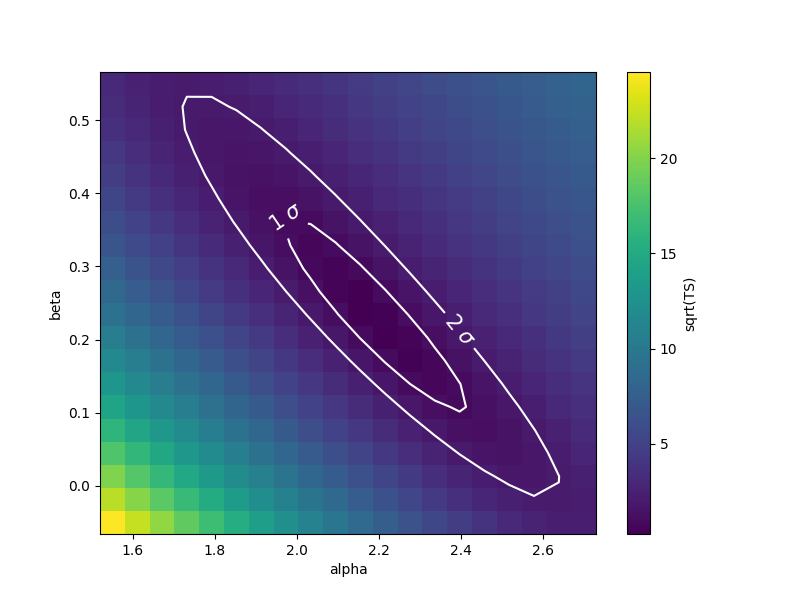
Note that, if computed with reoptimize=True, this plot would be
completely consistent with the third panel of the plot produced with
stat_contour (try!).
Finally, it is always remember that confidence contours are approximations. In particular, when the parameter range boundaries are close to the contours lines, it is expected that the statistical meaning of the contours is not well defined. That’s why we advise to always choose a parameter space that contains the contours you’re interested in.
Total running time of the script: ( 0 minutes 16.914 seconds)