This is a fixed-text formatted version of a Jupyter notebook
You may download all the notebooks in the documentation as a tar file.
Source files: flux_profiles.ipynb | flux_profiles.py
Flux Profile Estimation¶
This tutorial shows how to estimate flux profiles.
Prerequisites¶
Knowledge of 3D data reduction and datasets used in Gammapy, see for instance the first analysis tutorial.
Context¶
A useful tool to study and compare the saptial distribution of flux in images and data cubes is the measurement of flxu profiles. Flux profiles can show spatial correlations of gamma-ray data with e.g. gas maps or other type of gamma-ray data. Most commonly flux profiles are measured along some preferred coordinate axis, either radially distance from a source of interest, along longitude and latitude coordinate axes or along the path defined by two spatial coordinates.
Proposed Approach¶
Flux profile estimation essentially works by estimating flux points for a set of predefined spatially connected regions. For radial flux profiles the shape of the regions are annuli with a common center, for linear profiles it’s typically a rectangular shape.
We will work on a pre-computed MapDataset of Fermi-LAT data, use Region to define the structure of the bins of the flux profile and run the actually profile extraction using the FluxProfileEstimator
Setup and Imports¶
[1]:
%matplotlib inline
import matplotlib.pyplot as plt
from astropy import units as u
from astropy.coordinates import SkyCoord
import numpy as np
[2]:
from gammapy.datasets import MapDataset
from gammapy.estimators import FluxProfileEstimator, FluxPoints
from gammapy.maps import RegionGeom
from gammapy.utils.regions import (
make_concentric_annulus_sky_regions,
make_orthogonal_rectangle_sky_regions,
)
from gammapy.modeling.models import PowerLawSpectralModel
Read and Introduce Data¶
[3]:
dataset = MapDataset.read(
"$GAMMAPY_DATA/fermi-3fhl-gc/fermi-3fhl-gc.fits.gz", name="fermi-dataset"
)
This is what the counts image we will work with looks like:
[4]:
counts_image = dataset.counts.sum_over_axes()
counts_image.smooth("0.1 deg").plot(stretch="sqrt");
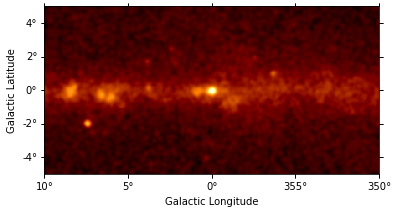

There are 400x200 pixels in the dataset and 11 energy bins between 10 GeV and 2 TeV:
[5]:
print(dataset.counts)
WcsNDMap
geom : WcsGeom
axes : ['lon', 'lat', 'energy']
shape : (400, 200, 11)
ndim : 3
unit :
dtype : >i4
Profile Estimation¶
Configuration¶
We start by defining a list of spatially connected regions along the galactic longitude axis. For this there is a helper function make_orthogonal_rectangle_sky_regions. The individual region bins for the profile have a height of 3 deg and in total there are 31 bins. The starts from lon = 10 deg tand goes to lon = 350 deg. In addition we have to specify the wcs to take into account possible projections effects on the region definition:
[6]:
regions = make_orthogonal_rectangle_sky_regions(
start_pos=SkyCoord("10d", "0d", frame="galactic"),
end_pos=SkyCoord("350d", "0d", frame="galactic"),
wcs=counts_image.geom.wcs,
height="3 deg",
nbin=51,
)
We can use the RegionGeom object to illustrate the regions on top of the counts image:
[7]:
geom = RegionGeom.create(region=regions)
ax = counts_image.smooth("0.1 deg").plot(stretch="sqrt")
geom.plot_region(ax=ax, color="w");
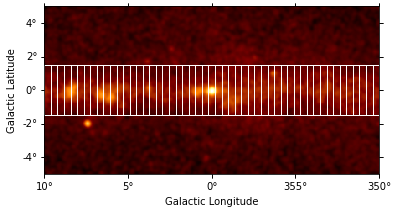
Next we create the FluxProfileEstimator. For the estimation of the flux profile we assume a spectral model with a power-law shape and an index of 2.3
[8]:
flux_profile_estimator = FluxProfileEstimator(
regions=regions,
spectrum=PowerLawSpectralModel(index=2.3),
energy_edges=[10, 2000] * u.GeV,
selection_optional=["ul"],
)
We can see the full configuration by printing the estimator object:
[9]:
print(flux_profile_estimator)
FluxProfileEstimator
--------------------
energy_edges : [ 10. 2000.] GeV
fit : <gammapy.modeling.fit.Fit object at 0x13f9adf40>
n_sigma : 1
n_sigma_ul : 2
norm_max : 5
norm_min : 0.2
norm_n_values : 11
norm_values : None
null_value : 0
reoptimize : False
selection_optional : ['ul']
source : 0
spectrum : PowerLawSpectralModel
sum_over_energy_groups : False
Run Estimation¶
Now we can run the profile estimation and explore the results:
[10]:
%%time
profile = flux_profile_estimator.run(datasets=dataset)
CPU times: user 6.87 s, sys: 121 ms, total: 6.99 s
Wall time: 6.76 s
[11]:
print(profile)
FluxPoints
----------
geom : RegionGeom
axes : ['lon', 'lat', 'energy', 'projected-distance']
shape : (1, 1, 1, 51)
quantities : ['norm', 'norm_err', 'norm_ul', 'ts', 'npred', 'npred_excess', 'stat', 'counts', 'success']
ref. model : pl
n_sigma : 1
n_sigma_ul : 2
sqrt_ts_threshold_ul : 2
sed type init : likelihood
We can see the flux profile is represented by a FluxPoints object with a projected-distance axis, which defines the main axis the flux profile is measured along. The lon and lat axes can be ignored.
Plotting Results¶
Let us directly plot the result using FluxPoints.plot():
[12]:
ax = profile.plot(sed_type="dnde")
ax.set_yscale("linear")
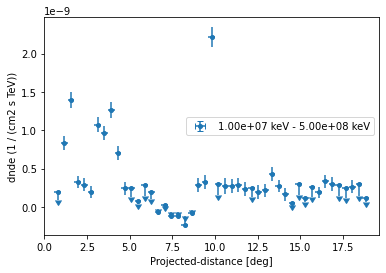
Based on the spectral model we specified above we can also plot in any other sed type, e.g. energy flux and define a different threshold when to plot upper limits:
[13]:
profile.sqrt_ts_threshold_ul = 2
ax = profile.plot(sed_type="eflux")
ax.set_yscale("linear")
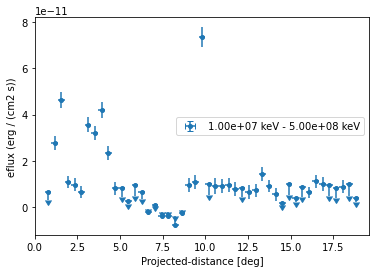
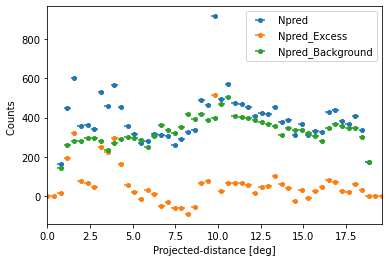
We can also plot any other quantity of interest, that is defined on the FluxPoints result object. E.g. the predicted total counts, background counts and excess counts:
[14]:
quantities = ["npred", "npred_excess", "npred_background"]
for quantity in quantities:
profile[quantity].plot(label=quantity.title())
plt.ylabel("Counts ")
[14]:
Text(0, 0.5, 'Counts ')
Serialisation and I/O¶
The profile can be serialised using FluxPoints.write(), given a specific format:
[15]:
profile.write(
filename="flux_profile_fermi.fits",
format="profile",
overwrite=True,
sed_type="dnde",
)
[16]:
profile_new = FluxPoints.read(
filename="flux_profile_fermi.fits", format="profile"
)
No reference model set for FluxMaps. Assuming point source with E^-2 spectrum.
[17]:
ax = profile_new.plot()
ax.set_yscale("linear")
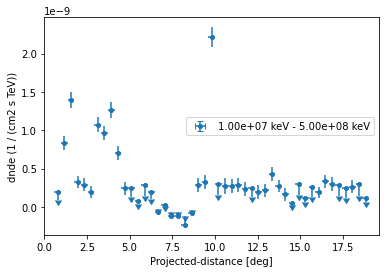
The profile can be serialised to a ~astropy.table.Table object using:
[18]:
table = profile.to_table(format="profile", formatted=True)
table
[18]:
| x_min | x_max | x_ref | e_ref [1] | e_min [1] | e_max [1] | ref_dnde [1] | ref_flux [1] | ref_eflux [1] | norm [1] | norm_err [1] | norm_ul [1] | ts [1] | sqrt_ts [1] | npred [1,1] | npred_excess [1,1] | stat [1] | is_ul [1] | counts [1,1] | success [1] |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| deg | deg | deg | keV | keV | keV | 1 / (cm2 s TeV) | keV / (cm2 s TeV) | keV2 / (cm2 s TeV) | |||||||||||
| float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float64 | float32 | float64 | bool | float64 | bool |
| -0.1960784313725492 | 0.1960784313725492 | 0.0 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | nan | nan | nan | nan | nan | nan | 0.0 | 0.000 | False | 0.0 | False |
| 0.1960784313725492 | 0.5882352941176467 | 0.392156862745098 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | nan | nan | nan | nan | nan | nan | 0.0 | 0.000 | False | 0.0 | False |
| 0.5882352941176467 | 0.9803921568627443 | 0.7843137254901955 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.172 | 0.124 | 0.434 | 2.059 | 1.435 | 162.23048986115955 | 17.448376 | -818.848 | True | 163.0 | True |
| 0.9803921568627443 | 1.3725490196078436 | 1.176470588235294 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 1.894 | 0.207 | 2.322 | 120.591 | 10.981 | 450.20181889217974 | 192.60909 | -3013.446 | False | 448.0 | True |
| 1.3725490196078436 | 1.764705882352942 | 1.5686274509803928 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 3.146 | 0.240 | 3.639 | 280.648 | 16.753 | 601.6822045016994 | 320.15994 | -4348.966 | False | 599.0 | True |
| 1.764705882352942 | 2.15686274509804 | 1.960784313725491 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.736 | 0.183 | 1.116 | 18.991 | 4.358 | 357.6650304886396 | 74.95968 | -2218.623 | False | 354.0 | True |
| 2.15686274509804 | 2.549019607843138 | 2.3529411764705888 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.656 | 0.186 | 1.042 | 14.214 | 3.770 | 363.83338136609376 | 66.89659 | -2452.717 | False | 367.0 | True |
| 2.549019607843138 | 2.941176470588236 | 2.745098039215687 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.450 | 0.179 | 0.821 | 7.011 | 2.648 | 339.4818273314676 | 45.943058 | -2174.005 | False | 339.0 | True |
| 2.941176470588236 | 3.3333333333333344 | 3.137254901960785 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 2.421 | 0.225 | 2.885 | 174.146 | 13.196 | 528.0599574489612 | 247.276 | -3887.531 | False | 531.0 | True |
| 3.3333333333333344 | 3.725490196078432 | 3.529411764705883 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 2.190 | 0.209 | 2.621 | 169.698 | 13.027 | 457.3561643542095 | 223.84384 | -3183.973 | False | 458.0 | True |
| 3.725490196078432 | 4.117647058823529 | 3.9215686274509807 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 2.868 | 0.232 | 3.345 | 243.821 | 15.615 | 564.4775740614491 | 293.2505 | -4201.048 | False | 566.0 | True |
| 4.117647058823529 | 4.509803921568627 | 4.313725490196078 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 1.599 | 0.205 | 2.023 | 81.901 | 9.050 | 451.76273200537855 | 163.63295 | -3040.782 | False | 448.0 | True |
| 4.509803921568627 | 4.901960784313727 | 4.7058823529411775 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.550 | 0.182 | 0.928 | 10.243 | 3.201 | 356.28517250965956 | 56.336945 | -2313.852 | False | 356.0 | True |
| 4.901960784313727 | 5.294117647058824 | 5.098039215686276 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.210 | 0.171 | 0.566 | 1.586 | 1.259 | 315.49805616415784 | 21.543198 | -1979.035 | True | 315.0 | True |
| 5.294117647058824 | 5.686274509803923 | 5.4901960784313735 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | -0.147 | 0.159 | 0.184 | 0.824 | -0.908 | 271.28752712430287 | -15.065349 | -1643.463 | True | 272.0 | True |
| 5.686274509803923 | 6.07843137254902 | 5.882352941176471 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.302 | 0.162 | 0.640 | 3.766 | 1.941 | 282.32873883100837 | 31.023876 | -1692.050 | True | 282.0 | True |
| 6.07843137254902 | 6.470588235294118 | 6.274509803921569 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.093 | 0.172 | 0.451 | 0.301 | 0.549 | 317.7330763355026 | 9.594112 | -2034.334 | True | 319.0 | True |
| 6.470588235294118 | 6.862745098039216 | 6.666666666666667 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | -0.500 | 0.170 | -0.147 | 7.804 | -2.794 | 311.3167047326352 | -51.37142 | -1931.193 | True | 311.0 | True |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 12.352941176470619 | 12.745098039215696 | 12.549019607843157 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.441 | 0.197 | 0.848 | 5.381 | 2.320 | 424.9721062687647 | 45.757168 | -2877.351 | False | 425.0 | True |
| 12.745098039215696 | 13.137254901960773 | 12.941176470588236 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.492 | 0.194 | 0.893 | 7.018 | 2.649 | 416.7588470906105 | 51.088707 | -2705.136 | False | 412.0 | True |
| 13.137254901960773 | 13.529411764705875 | 13.333333333333325 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.959 | 0.205 | 1.382 | 25.879 | 5.087 | 455.22507305193835 | 99.72924 | -3216.032 | False | 458.0 | True |
| 13.529411764705875 | 13.921568627451002 | 13.72549019607844 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.611 | 0.184 | 0.993 | 12.465 | 3.531 | 376.66733293729885 | 63.567978 | -2403.516 | False | 373.0 | True |
| 13.921568627451002 | 14.31372549019608 | 14.117647058823541 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.377 | 0.187 | 0.763 | 4.407 | 2.099 | 385.1566764851361 | 39.272778 | -2526.278 | False | 384.0 | True |
| 14.31372549019608 | 14.705882352941185 | 14.509803921568633 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | -0.244 | 0.168 | 0.103 | 2.022 | -1.422 | 313.3736443463295 | -25.44988 | -1936.261 | True | 313.0 | True |
| 14.705882352941185 | 15.098039215686311 | 14.901960784313747 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.307 | 0.179 | 0.679 | 3.133 | 1.770 | 366.322547445814 | 32.01158 | -2178.012 | True | 357.0 | True |
| 15.098039215686311 | 15.490196078431389 | 15.294117647058851 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | -0.083 | 0.167 | 0.265 | 0.240 | -0.490 | 313.4101919680869 | -8.618814 | -1914.901 | True | 311.0 | True |
| 15.490196078431389 | 15.882352941176466 | 15.686274509803926 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.228 | 0.172 | 0.584 | 1.863 | 1.365 | 330.89113194006126 | 23.80037 | -2016.119 | True | 327.0 | True |
| 15.882352941176466 | 16.274509803921596 | 16.07843137254903 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.427 | 0.170 | 0.780 | 6.979 | 2.642 | 324.69019708074285 | 44.554733 | -1957.576 | False | 321.0 | True |
| 16.274509803921596 | 16.666666666666696 | 16.470588235294144 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.760 | 0.195 | 1.163 | 17.553 | 4.190 | 428.005206597181 | 79.45124 | -2767.621 | False | 421.0 | True |
| 16.666666666666696 | 17.058823529411775 | 16.862745098039234 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.675 | 0.197 | 1.082 | 13.277 | 3.644 | 437.13415063137586 | 70.588745 | -2830.574 | False | 431.0 | True |
| 17.058823529411775 | 17.4509803921569 | 17.254901960784338 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.256 | 0.183 | 0.634 | 2.057 | 1.434 | 381.24750962187824 | 26.727816 | -2356.744 | True | 374.0 | True |
| 17.4509803921569 | 17.843137254902004 | 17.647058823529452 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.174 | 0.182 | 0.551 | 0.951 | 0.975 | 366.45334911783 | 18.242867 | -2463.364 | True | 370.0 | True |
| 17.843137254902004 | 18.235294117647083 | 18.039215686274545 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.593 | 0.192 | 0.990 | 10.705 | 3.272 | 408.05504964497743 | 62.12828 | -2803.387 | False | 410.0 | True |
| 18.235294117647083 | 18.627450980392158 | 18.43137254901962 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.306 | 0.173 | 0.666 | 3.342 | 1.828 | 334.14141424669504 | 32.074398 | -2186.572 | True | 336.0 | True |
| 18.627450980392158 | 19.01960784313726 | 18.82352941176471 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | 0.010 | 0.124 | 0.269 | 0.005 | 0.071 | 171.8115649139593 | 0.9959075 | -877.032 | True | 172.0 | True |
| 19.01960784313726 | 19.41176470588239 | 19.215686274509824 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | nan | nan | nan | nan | nan | nan | 0.0 | 0.000 | False | 0.0 | False |
| 19.41176470588239 | 19.803921568627494 | 19.607843137254942 | 70710678.119 | 10000000.000 | 500000000.000 | 4.428e-10 | 3.043e-01 | 9.166e+06 | nan | nan | nan | nan | nan | nan | 0.0 | 0.000 | False | 0.0 | False |
No we can also estimate a radial profile starting from the Galactic center:
[19]:
regions = make_concentric_annulus_sky_regions(
center=SkyCoord("0d", "0d", frame="galactic"),
radius_max="1.5 deg",
nbin=11,
)
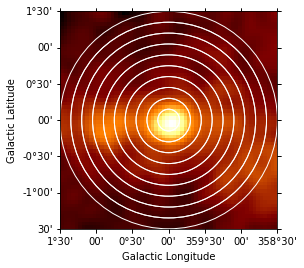
Again we first illustrate the regions:
[20]:
geom = RegionGeom.create(region=regions)
gc_image = counts_image.cutout(
position=SkyCoord("0d", "0d", frame="galactic"), width=3 * u.deg
)
ax = gc_image.smooth("0.1 deg").plot(stretch="sqrt")
geom.plot_region(ax=ax, color="w");
/Users/adonath/software/mambaforge/envs/gammapy-dev/lib/python3.9/site-packages/scipy/optimize/_numdiff.py:579: RuntimeWarning: invalid value encountered in true_divide
J_transposed[i] = df / dx
This time we define two energy bins and include the fit statistic profile in the computation:
[21]:
flux_profile_estimator = FluxProfileEstimator(
regions=regions,
spectrum=PowerLawSpectralModel(index=2.3),
energy_edges=[10, 100, 2000] * u.GeV,
selection_optional=["ul", "scan"],
norm_values=np.linspace(-1, 5, 11),
)
[22]:
profile = flux_profile_estimator.run(datasets=dataset)
We can directly plot the result:
[23]:
ax = profile.plot(axis_name="projected-distance", sed_type="flux")
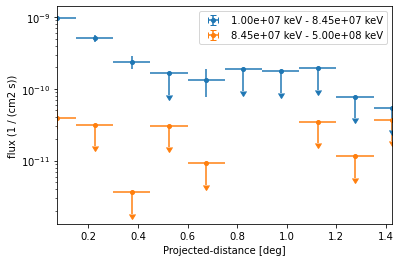
However because of the powerlaw spectrum the flux at high energies is much lower. To extract the profile at high energies only we can use:
[24]:
profile_high = profile.slice_by_idx({"energy": slice(1, 2)})
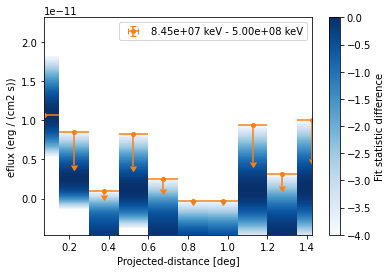
And now plot the points together with the likelihood profiles:
[25]:
ax = profile_high.plot(sed_type="eflux", color="tab:orange")
profile_high.plot_ts_profiles(ax=ax, sed_type="eflux")
ax.set_yscale("linear")
[ ]:
